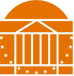
Lipoprotein Signal Peptidase (LspA)
With the rapid growth of antibiotic resistance, new drugs are needed which target novel pathways. The lipoprotein processing pathway is a novel pathway for antibiotic drug targeting as the enzymes involved, Lgt, LspA, and Lnt, are essential in some organisms including E. coli, S. enterica, M. tuberculosis, and S. coelicolor, and have no mammalian homologs. Lipoprotein signal peptidase (LspA) is an aspartyl protease that carries out the second step in the lipoprotein processing pathway - cleaving the transmembrane helical signal peptide of lipoproteins after lipidation by Lgt.

The crystal structure of LspA has been determined in complex with the antibiotic globomycin. However, the apo and lipoprotein substrate bound structures of LspA remain elusive.
We are investigating the conformational dynamics of LspA using electron paramagnetic resonance (EPR) studies and molecular dynamics (MD) simulations. This hybrid approach allows for visualization of structures consistent with experimental EPR restraints.